DNAmoreDB is designed to contain comprehensive information of all DNAzymes that have been isolated by in vitro selection and in vitro evolution. In addition to sequence information, you will find links to the original publications, reaction conditions, reaction substrates, metal-ion requirements, and structural information whenever available. The inclusion of a BLASTN search facilitates the comparison of newly selected sequences with already existing ones; while the advanced search can retrieve specific content of the database with the use of keywords.
The application programming interface (API) gives the advanced user the ability to retrieve information about DNAzymes through simple URL schemes. The API offers the possibility for developers to connect their applications directly to DNAmoreDB, they can do so by defining the methods to request data from the database. To understand how the DNAmoreDB API system can be accessed please visit the API page.
Although DNAmoreDB is updated monthly, we encourage authors and users to submit their data by filling the form under the Submit tab.
To help you navigate DNAmoreDB we have created this page. Some of the descriptors that we use are explained here in detail (e.g. Reacting groups, Sequence description, etc).
There is no naming convention for DNAzymes, instead, each laboratory uses its own. Some of the names make reference to the in vitro selection round from which the DNAzyme was isolated, to the in vitro selection’s name (experiment’s name or reference), and to the clone number. For instance, 9DB1 would refer to “round 9 of the DB selection, clone 1”. Other names are given according to a classification made by the authors based on sequence similarity, for example, “Class 1”. This kind of name may not be as unique as the former one, and therefore it is necessary, in order to identify the DNAzyme unambiguously, to mention additional attributes such as the catalytic activity, i.e. RNA-cleaving Class 1 DNAzyme, or DNA-capping Class 1 DNAzyme.
The catalytic region of a DNAzyme corresponds to the part of the DNAzyme that was allowed to evolve during the in vitro selection experiment. Normally, it is flanked by constant regions that are primer binding sites. The length of the catalytic region is decided upon before the in vitro selection experiment takes place and varies, usually, between 20 and 80 nucleotides (figure 1). Shorter catalytic regions allow for better coverage of the sequence space, but at the same time are expected to be able to form less complex catalytic centers; the opposite is true for longer catalytic regions. The fact that we refer to this region of the DNAzyme as “catalytic”, does not imply that all the nucleotides in it are necessary for the DNAzyme’s catalytic activity. Combinatorial methods, probing, and mutagenesis experiments coupled to activity assays, as well as structural studies, allow to discern which nucleotides are essential for the DNAzyme to catalyze a reaction, or to adopt the fold that ultimately will make catalysis possible.
In addition, you will realize that there is no standard way for reporting DNAzymes' sequences, and each laboratory generally has a preference on how to do so. For instance, some choose to report only the catalytic region (Nx) of the isolated DNAzymes, while others report the sequence of the "whole" sequenced DNA, meaning that constant regions and the catalytic region are reported, often along with the substrate (if it was ligated to the pool). In DNAmoreDB, when the length of the annotated DNAzyme sequence matches that of the pool (Nx) plus/minus a couple of nucleotides, the annotated sequence corresponds to that of the catalytic region. Instead, if the length of the annotated DNAzyme is longer (or much longer) than the pool, the sequence has been annotated along with the constant regions (and at times even with the substrate).
In the context of secondary structure prediction of DNAzymes of course one needs to take into account not only the catalytic region of the DNAzyme, but also the substrate recognition domains (also referred to as constant regions or binding arms) and the substrate itself.
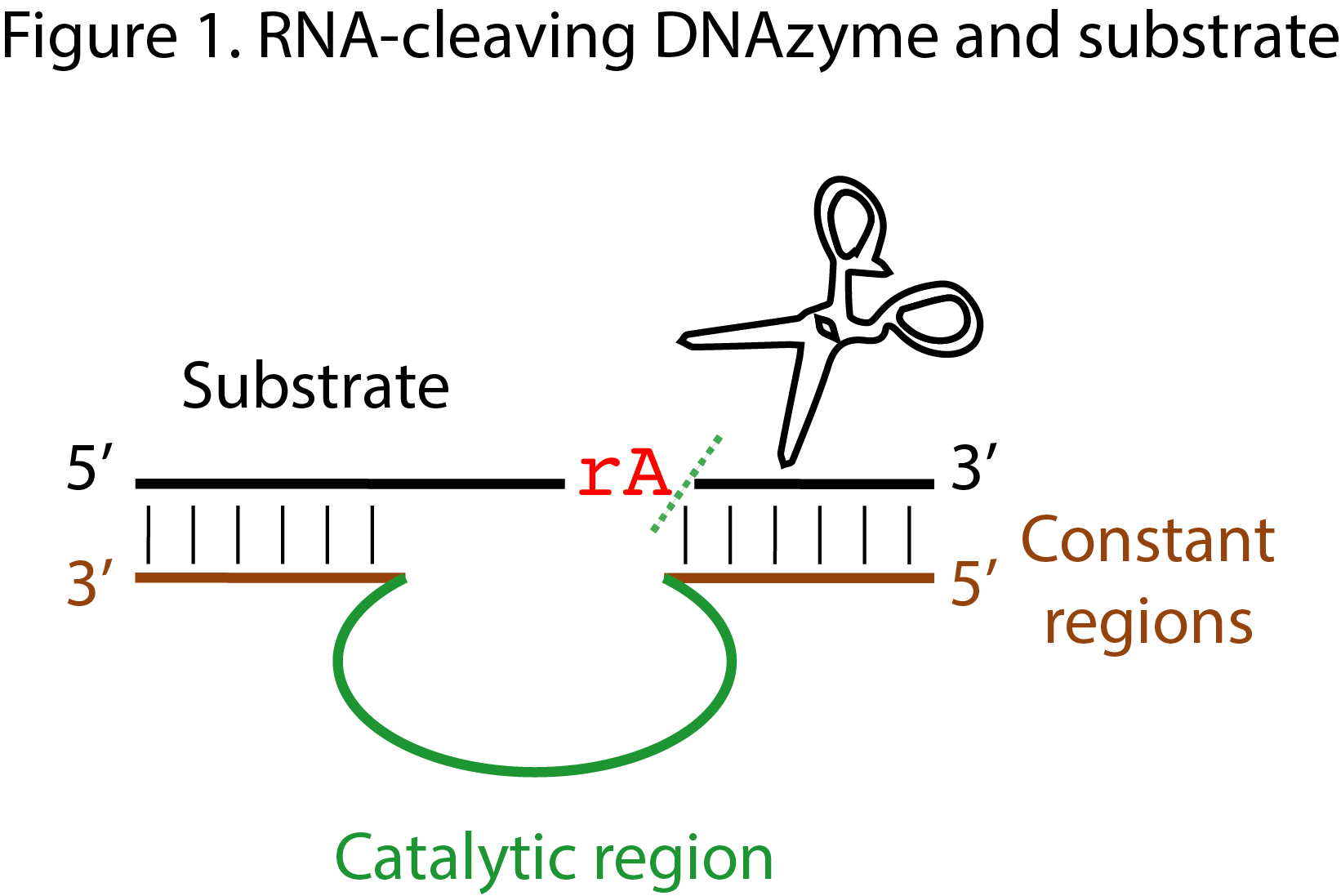
This is the reaction the selected DNAzyme catalyzes in the reaction buffer, in the presence of its substrates and cofactors. Note that some DNAzymes are selected to catalyze the reaction in cis, while others are selected to catalyze the reaction in trans. Often, though, authors report trans-acting DNAzymes although during the original selection the isolated DNAzymes acted in cis. For more information about the reactions, please visit the Reactions page or simply click on the name of the reaction.
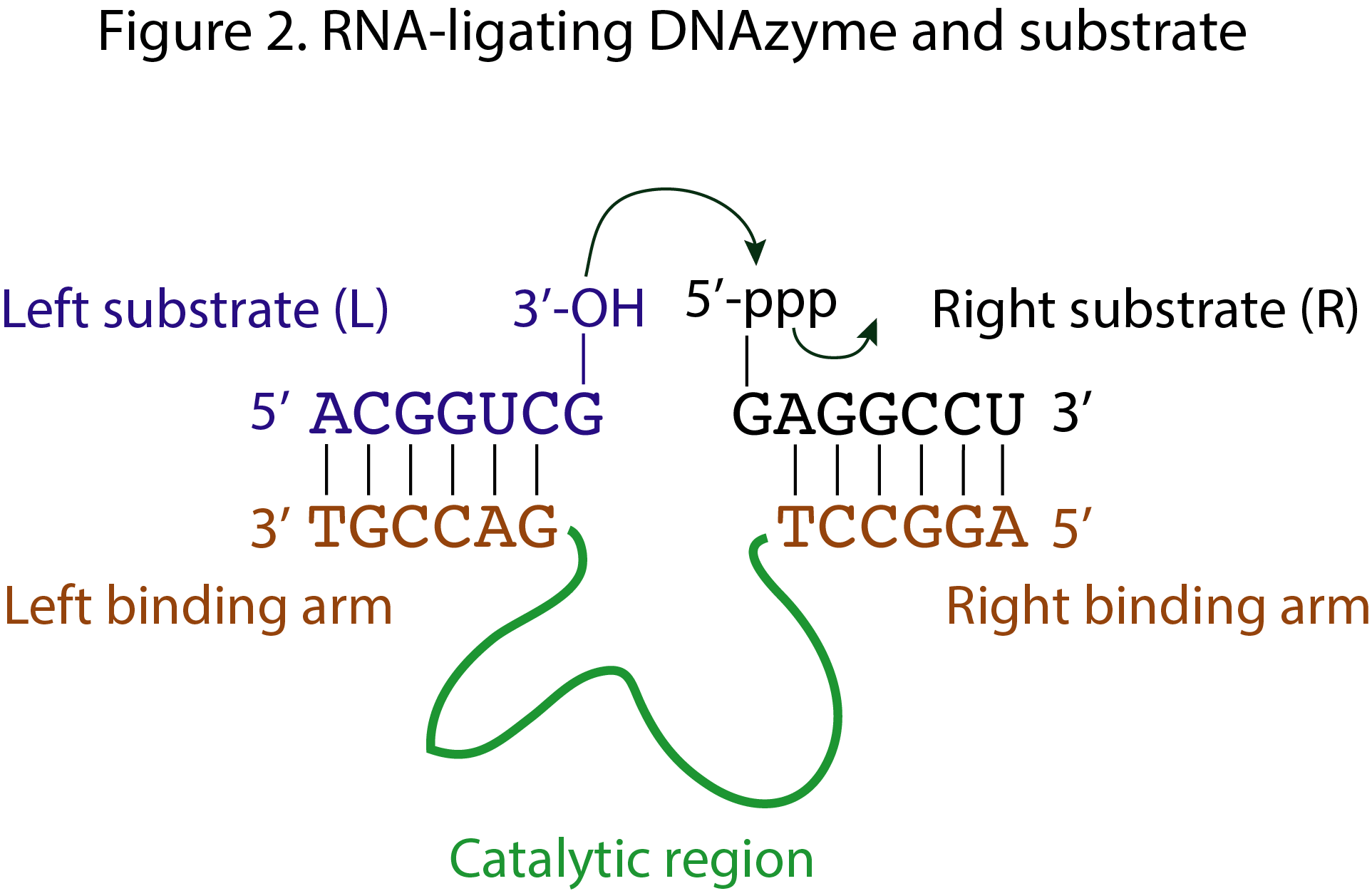
Nucleic acid substrates of DNAzymes are recognized by means of Watson-Crick base pairing. When there are two substrates, L and R, they hybridize to the left and right “arms” of the DNAzyme (figure 2). When there is a unique substrate, we refer to it as S and the binding arms will be referred to as left and right depending on the position in the substrate where the chemical transformation occurs. If the substrates carry any modification, it will be indicated as X within the substrate sequence, and later explained, i.e. entry 8TM8 uses two substrates, L and R, where the L substrate has sequence GGATAATACGACTCACTATX, with X being a flexible linker (hexaethylene glycol) and a tripeptide Cysteine-Tyrosine-Alanine (X: HEG-Cys-Tyr-Ala).
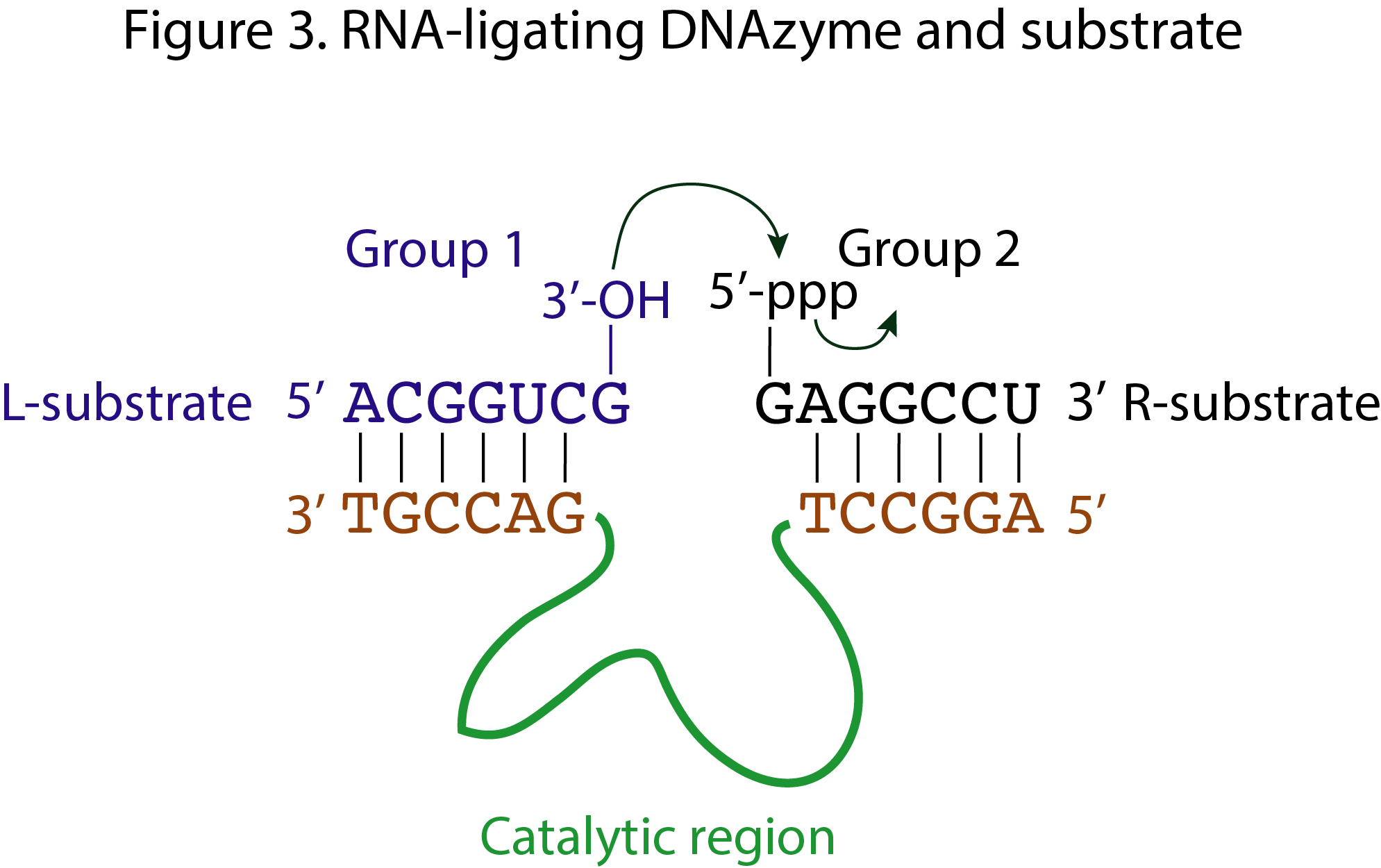
These are the functional groups or residues taking part in the reaction. While Group 1 refers to the 3’ end of the 5’ substrate, L, or left substrate; Group 2 refers to the 5’ end of the 3’ substrate, R, or right substrate (figure 3). Of course, it is also possible that the two reacting groups are found within the same substrate (S).
Although there are instances of metal ion-independent and cofactor-independent DNAzymes, most of them require the presence of divalent metal ions for their activity. However, their roles, whether catalytic or structural, are difficult to define without structural information.
This field simply indicates the length of the random sequence region of the DNA pool that was subjected to the in vitro selection experiment. It is referred to as N followed by the length of the random region, e.g. N40. The catalytic region of the DNAzymes isolated from the in vitro selection experiment will normally have the same length, although at times there can be insertions or deletions caused by Taq polymerase during the selection process.
These are incubation conditions during the key selection step of the in vitro selection experiment. For information about the incubation time and incubation temperature please refer to the main publication. The metal ions and cofactors are listed within the buffer conditions if they were present during the in vitro selection key selection step. However, at times, you may encounter DNAzymes that can utilize several metal ions, some of which were not listed in the buffer composition. These “additional” metal ions in which presence the DNAzyme can catalyze the desired reaction are found through subsequent experiments.
For the best candidate from each in vitro selection, as well as for those for which this information was available, we have annotated the rate constant and/or final yields. These refer to a given reaction condition, a given substrate and a given setup (cis or trans). However, bear in mind that several rate constants and yields may be available for a DNAzyme if it has been tested or validated under different conditions of pH, temperature, metal ions, cofactors, point mutations, etc. Therefore, for further details on a given DNAzyme's kinetic characterization you should refer to the publication in which it was reported. The sequences for which reaction rate and/or final yield are available appear in bold characters in the DNAzymes page. This information provides guidelines to help you decide which sequence to pick for further studies or for a desired application since not all the sequences reported after an in vitro selection experiment are characterized or validated to the same extent, and many, in fact, may not even be active.
"Reported in" is the publication where a given DNAzyme was first reported.
Related publications are those that report further results about a given DNAzyme, or closely related ones. The related publications mention specifically within the main text of the article the name of the DNAzyme in question. If your publication refers to any of the DNAzymes in our database and should be included as a related publication for a given DNAzyme, please write to us using the form under the Submit tab.